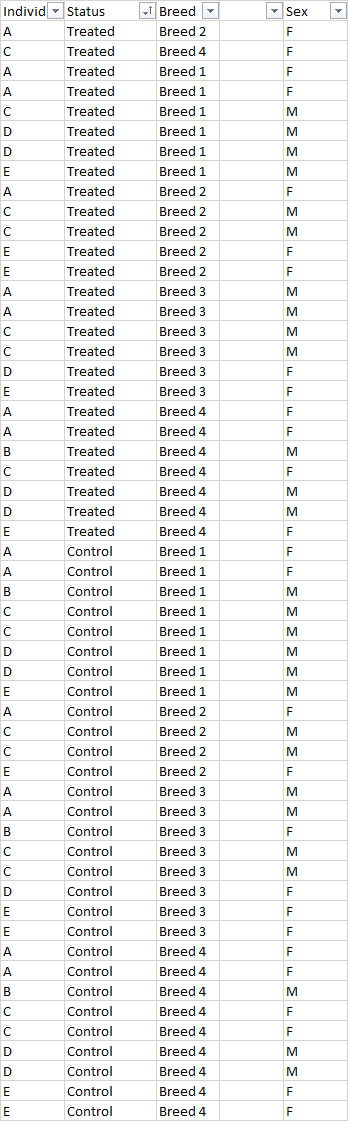
Dear All, I have a tricky question regarding the use of EdgeR in R to run differential expression analyses of the following experiment: it is a multifactorial experiment with around 55 sample designed as follow: Conditions = 2 (treatment & Control) Breed = 4 breeds Sex of animals = 2 (male and Female)
Within each breed, I have samples from 5 individuals (A, B, C, D and E)
I have two main questions:
I know that I can adjust for additional variables upon DE analyses, but When running an nMDS looking at how treated subjects would seprats from control, I can not really visualize this, while adjusting for differences in gender or breed using the nMDS approach ... Are there any idea or technique so that when visualizing the clustering of treatment vs Ctr samples to adjust for other covariate like sex or breed in here ? I need to do this because if I see clustering, I am afriad that this could be due to sex-specific effect, and not necessarily a treatment effect.
If I am going to calculate DE genes between treatmeent & Ctr, I think I need to adjust for 1) base line differences between individuals (as this is a paired design because the same animals were blood treated and as a Control), 2) sex differences 3)Breed.
Could you advice, which of the designs I should follow from the updated guidlines of edgeR (https://www.bioconductor.org/packages/devel/bioc/vignettes/edgeR/inst/doc/edgeRUsersGuide.pdf) that could match my experiment ? I though about the one in section 3.4.1, but with more factors and adjusting for them.
Thanks a lot for any comments. .. Code should be placed in three backticks as shown below


Cross-posted to Biostars https://www.biostars.org/p/9520241/
The analysis depends on what questions you want to ask.
Let us assume I want to test treatement Vs control Overall : so using nMDS for that (the first question), how can I adjust for sex/breed variation. Because I assume if one see sepration (or even not), this could be due to gende difference or breed differences between control or treatment subjects rather than the treatment itself
If I am going to run DE looking at differences between treatment/control, I need also to adjust for that.
Could you advice on which example given in the tutorial (link is above) fits the case when I need to compare between treatment and control overall ? taking a note that this is a paired design, so many of the treated subjects were the same used as controls. Thanks